Home¶
ggCaller is a novel bacterial gene annotation and pangenome analysis tool, designed to enable fast, accurate analysis of large single-species genome datasets.
ggCaller traverses de Bruijn graphs (DBGs) built by Bifrost, using temporal convolutional networks from Balrog for gene filtering and Panaroo for pangenome analysis and quality control.
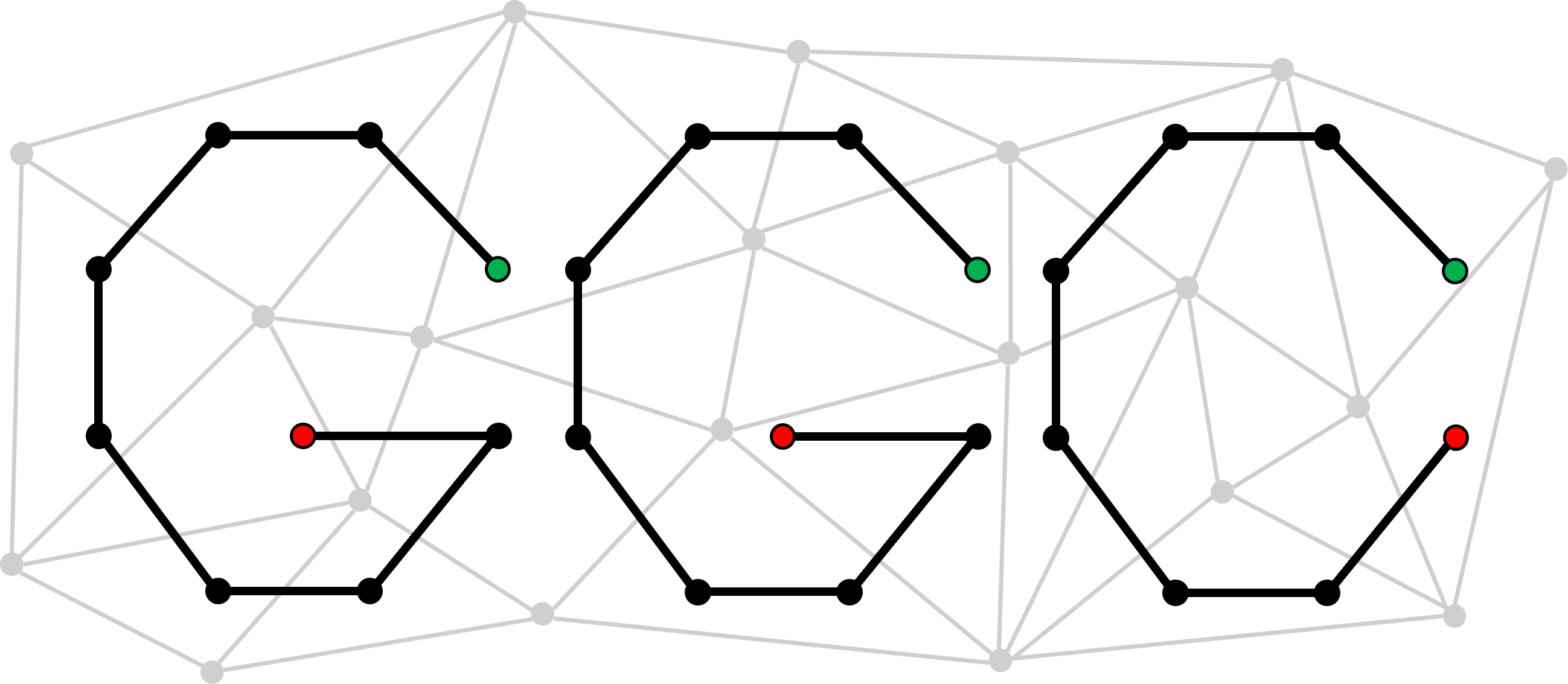
Why ggCaller?¶
ggCaller uses population-frequency information at several stages of gene annotation and pangenome analysis. This has several benefits:
Consistent identification of start and stop codons across orthologs, improving clustering accuracy.
Reduced gene-annotation sensitivity to assembly fragmentation.
Reduced runtime verses existing gene-annotation and pangenome analysis workflows.
One-line command from fasta -> gene annotations, gene frequency matrices, clusters of orthologous genes (COGs), core genome/pangenome alignments, phylogenetic trees, small/structural variants and more!
Annotated DBG-querying for functional PanGenome-Wide Association Studies (PGWAS), compatible with results from Pyseer.
For the impatient¶
See Quickstart to get ggCaller up and running quickly.
Everyone else¶
We recommend starting with Installation to ensure things are installed correctly, followed by Usage to get an overview of the commands, and finally Tutorial for a step-by-step walkthrough.